Broad Institute of MIT and Harvard
22 Oct 2025
TOOLUNIVERSE establishes an open-source ecosystem that standardizes AI-tool interaction and empowers AI scientists to autonomously discover, create, optimize, and compose scientific tools. The platform successfully demonstrated its utility in a therapeutic discovery case study, identifying a novel drug candidate for hypercholesterolemia with validated properties.
14 Mar 2025
Researchers from Harvard Medical School and partners develop TxAgent, an AI agent that combines large language models with 211 biomedical tools for therapeutic reasoning, outperforming GPT-4 by up to 25.8% on drug selection tasks while providing evidence-grounded treatment recommendations through multi-step reasoning traces.
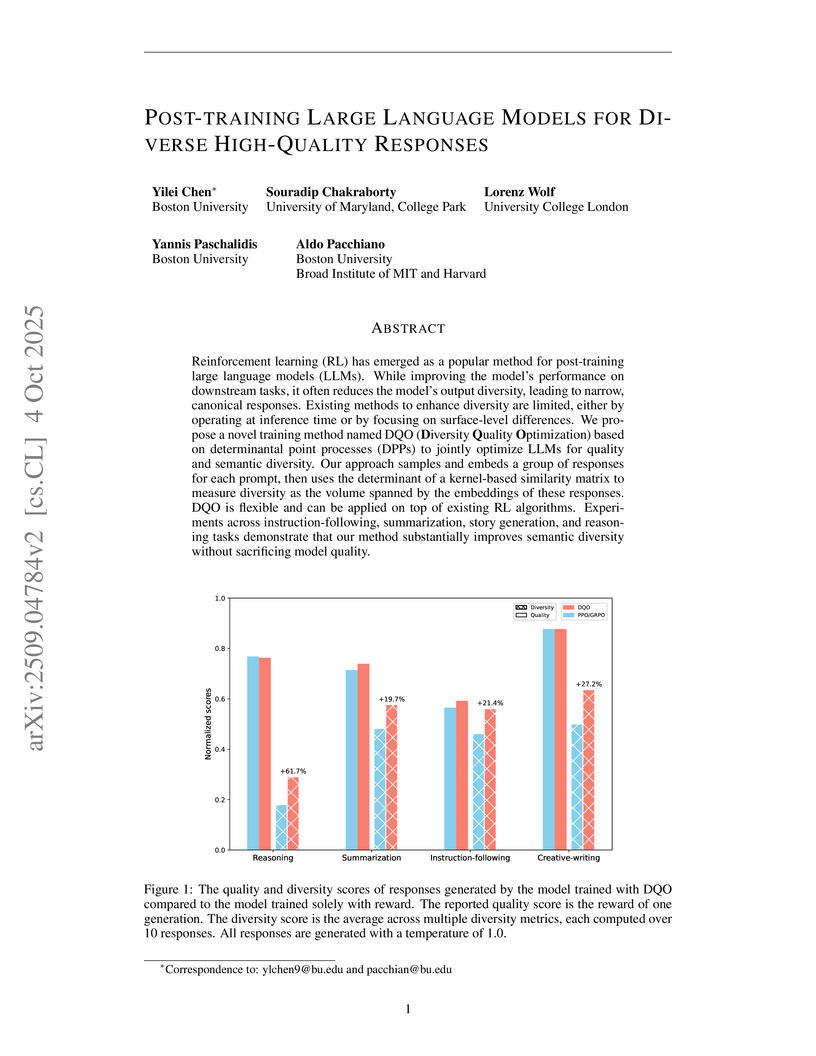
Reinforcement learning (RL) has emerged as a popular method for post-training large language models (LLMs). While improving the model's performance on downstream tasks, it often reduces the model's output diversity, leading to narrow, canonical responses. Existing methods to enhance diversity are limited, either by operating at inference time or by focusing on surface-level differences. We propose a novel training method named DQO (Diversity Quality Optimization) based on determinantal point processes (DPPs) to jointly optimize LLMs for quality and semantic diversity. Our approach samples and embeds a group of responses for each prompt, then uses the determinant of a kernel-based similarity matrix to measure diversity as the volume spanned by the embeddings of these responses. DQO is flexible and can be applied on top of existing RL algorithms. Experiments across instruction-following, summarization, story generation, and reasoning tasks demonstrate that our method substantially improves semantic diversity without sacrificing model quality.
STFlow is a computational framework that predicts spatially-resolved gene expression directly from standard histology whole-slide images, eliminating the need for expensive wet-lab experiments. It leverages a flow matching approach to model joint gene expression distributions and achieved an 18% average relative improvement in prediction performance over pathology foundation models while demonstrating significantly enhanced computational efficiency.
Inference from tabular data, collections of continuous and categorical variables organized into matrices, is a foundation for modern technology and science. Yet, in contrast to the explosive changes in the rest of AI, the best practice for these predictive tasks has been relatively unchanged and is still primarily based on variations of Gradient Boosted Decision Trees (GBDTs). Very recently, there has been renewed interest in developing state-of-the-art methods for tabular data based on recent developments in neural networks and feature learning methods. In this work, we introduce xRFM, an algorithm that combines feature learning kernel machines with a tree structure to both adapt to the local structure of the data and scale to essentially unlimited amounts of training data.
We show that compared to 31 other methods, including recently introduced tabular foundation models (TabPFNv2) and GBDTs, xRFM achieves best performance across 100 regression datasets and is competitive to the best methods across 200 classification datasets outperforming GBDTs. Additionally, xRFM provides interpretability natively through the Average Gradient Outer Product.
 University of Washington
University of Washington University of Toronto
University of Toronto Harvard University
Harvard University Carnegie Mellon University
Carnegie Mellon University Stanford University
Stanford University University of California, San Diego
University of California, San Diego Google Research
Google Research Northwestern University
Northwestern University Columbia UniversityVector InstituteHarvard Medical School
Columbia UniversityVector InstituteHarvard Medical School EPFL
EPFL Lawrence Berkeley National Laboratory
Lawrence Berkeley National Laboratory Mohamed bin Zayed University of Artificial Intelligence
Mohamed bin Zayed University of Artificial Intelligence Technical University of Munich
Technical University of Munich KTH Royal Institute of TechnologyUniversity of California San FranciscoHoward Hughes Medical InstituteHelmholtz Center MunichGenentechBroad Institute of MIT and HarvardEuropean Molecular Biology LaboratoryChan Zuckerberg InitiativeChan Zuckerberg BiohubKempner Institute for the Study of Natural and Artificial IntelligenceCalico Life Sciences LLCArc InstituteBrotman Baty Institute for Precision MedicineGladstone Institute of Cardiovascular DiseaseEvolutionaryScale, PBCSchmidt FuturesCellarityNewLimit
KTH Royal Institute of TechnologyUniversity of California San FranciscoHoward Hughes Medical InstituteHelmholtz Center MunichGenentechBroad Institute of MIT and HarvardEuropean Molecular Biology LaboratoryChan Zuckerberg InitiativeChan Zuckerberg BiohubKempner Institute for the Study of Natural and Artificial IntelligenceCalico Life Sciences LLCArc InstituteBrotman Baty Institute for Precision MedicineGladstone Institute of Cardiovascular DiseaseEvolutionaryScale, PBCSchmidt FuturesCellarityNewLimitA collaboration of over 40 experts from diverse institutions, supported by the Chan Zuckerberg Initiative, outlines a comprehensive vision for the AI Virtual Cell (AIVC), a multi-scale, multi-modal neural network model to represent and simulate cellular behavior. This framework aims to integrate vast biological datasets with advanced AI to accelerate biomedical discovery and enable in silico experimentation.
 Stanford University
Stanford University University of Michigan
University of Michigan Cornell UniversityHarvard Medical School
Cornell UniversityHarvard Medical School The University of Hong Kong
The University of Hong Kong Rutgers University
Rutgers University Shandong UniversityUniversity of OttawaChinese Academy of Medical SciencesPeking University Sixth HospitalBroad Institute of MIT and HarvardChinese Academy of Medical Sciences and Peking Union Medical CollegeMassGeneral Brigham
Shandong UniversityUniversity of OttawaChinese Academy of Medical SciencesPeking University Sixth HospitalBroad Institute of MIT and HarvardChinese Academy of Medical Sciences and Peking Union Medical CollegeMassGeneral BrighamAIPatient, a simulated patient system, integrates a multi-agent large language model architecture with a knowledge graph built from real-world patient data. It demonstrates high accuracy in medical question-answering and superior fidelity, usability, and educational effectiveness compared to human-simulated patients in a user study.
Large language models (LLMs) hold great promise for medical applications and are evolving rapidly, with new models being released at an accelerated pace. However, benchmarking on large-scale real-world data such as electronic health records (EHRs) is critical, as clinical decisions are directly informed by these sources, yet current evaluations remain limited. Most existing benchmarks rely on medical exam-style questions or PubMed-derived text, failing to capture the complexity of real-world clinical data. Others focus narrowly on specific application scenarios, limiting their generalizability across broader clinical use. To address this gap, we present BRIDGE, a comprehensive multilingual benchmark comprising 87 tasks sourced from real-world clinical data sources across nine languages. It covers eight major task types spanning the entire continuum of patient care across six clinical stages and 20 representative applications, including triage and referral, consultation, information extraction, diagnosis, prognosis, and billing coding, and involves 14 clinical specialties. We systematically evaluated 95 LLMs (including DeepSeek-R1, GPT-4o, Gemini series, and Qwen3 series) under various inference strategies. Our results reveal substantial performance variation across model sizes, languages, natural language processing tasks, and clinical specialties. Notably, we demonstrate that open-source LLMs can achieve performance comparable to proprietary models, while medically fine-tuned LLMs based on older architectures often underperform versus updated general-purpose models. The BRIDGE and its corresponding leaderboard serve as a foundational resource and a unique reference for the development and evaluation of new LLMs in real-world clinical text understanding.
The BRIDGE leaderboard: this https URL
Researchers at MIT and UC San Diego characterize a fundamental mechanism of feature learning in deep fully connected networks: that networks up-weight features proportional to the average gradient outer product. Leveraging this insight, they introduce Recursive Feature Machines (RFMs), a new class of models that achieve state-of-the-art performance on tabular datasets with a significant reduction in computational cost, while also providing explanations for several deep learning phenomena.
MADRIGAL, a multimodal AI model developed by researchers including those from Harvard Medical School and AstraZeneca, predicts clinical outcomes of drug combinations using diverse preclinical data while robustly handling missing information. The model significantly improves prediction of adverse events and patient-specific treatment responses, demonstrating strong alignment with real-world clinical trial data and electronic health records.
27 Sep 2025
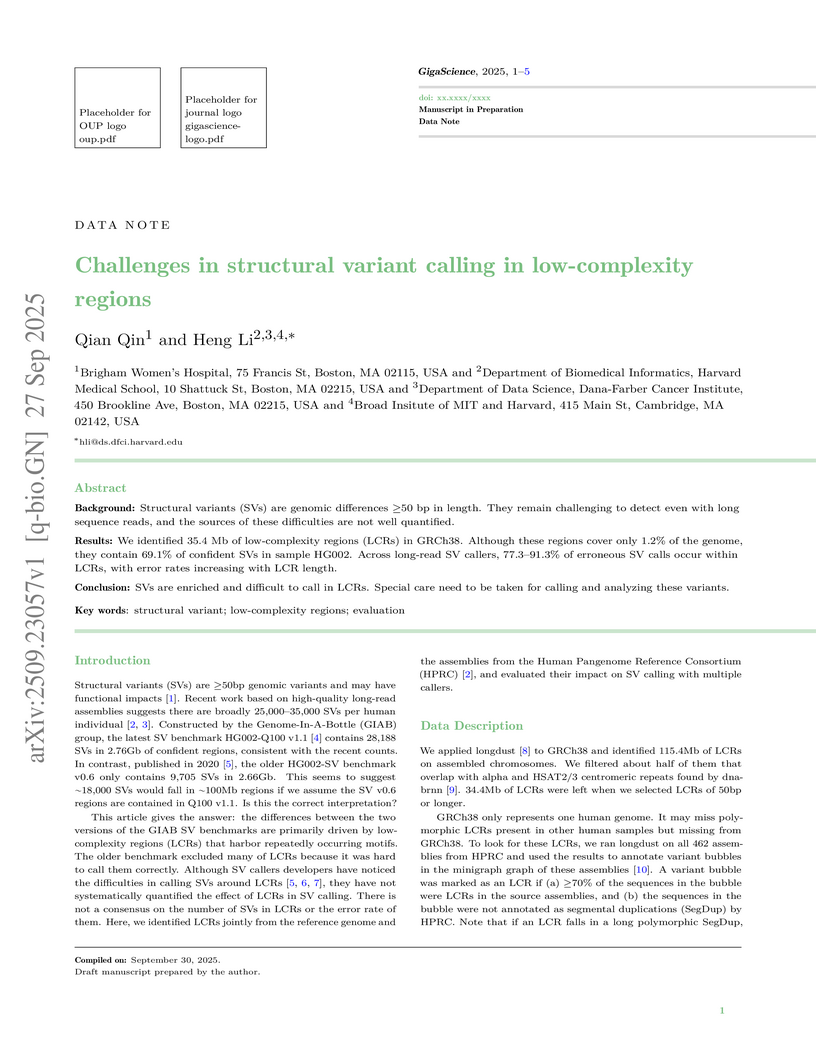
Background: Structural variants (SVs) are genomic differences ≥50 bp in length. They remain challenging to detect even with long sequence reads, and the sources of these difficulties are not well quantified.
Results: We identified 35.4 Mb of low-complexity regions (LCRs) in GRCh38. Although these regions cover only 1.2% of the genome, they contain 69.1% of confident SVs in sample HG002. Across long-read SV callers, 77.3-91.3% of erroneous SV calls occur within LCRs, with error rates increasing with LCR length.
Conclusion: SVs are enriched and difficult to call in LCRs. Special care need to be taken for calling and analyzing these variants.
Petrović et al. introduce CURLY FLOW MATCHING (CURLY-FM), a simulation-free machine learning framework that learns complex, non-gradient field dynamics by solving a Schrödinger bridge problem with a non-zero drift reference process. The method reconstructs periodic trajectories in diverse scientific domains like ocean currents and cell cycles, outperforming existing flow-matching and simulation-based techniques in efficiency and accuracy.
A perspective paper outlines a comprehensive vision for "AI scientists" as collaborative AI agents in biomedical discovery, proposing a modular architectural framework and a four-level autonomy taxonomy. This approach aims to accelerate research workflows, enable novel insights beyond current capabilities, and transform human-AI collaboration.
A digital twin is a virtual replica of a real-world physical phenomena that
uses mathematical modeling to characterize and simulate its defining features.
By constructing digital twins for disease processes, we can perform in-silico
simulations that mimic patients' health conditions and counterfactual outcomes
under hypothetical interventions in a virtual setting. This eliminates the need
for invasive procedures or uncertain treatment decisions. In this paper, we
propose a method to identify digital twin model parameters using only
noninvasive patient health data. We approach the digital twin modeling as a
composite inverse problem, and observe that its structure resembles pretraining
and finetuning in self-supervised learning (SSL). Leveraging this, we introduce
a physics-informed SSL algorithm that initially pretrains a neural network on
the pretext task of learning a differentiable simulator of a physiological
process. Subsequently, the model is trained to reconstruct physiological
measurements from noninvasive modalities while being constrained by the
physical equations learned in pretraining. We apply our method to identify
digital twins of cardiac hemodynamics using noninvasive echocardiogram videos,
and demonstrate its utility in unsupervised disease detection and in-silico
clinical trials.
As post hoc explanations are increasingly used to understand the behavior of graph neural networks (GNNs), it becomes crucial to evaluate the quality and reliability of GNN explanations. However, assessing the quality of GNN explanations is challenging as existing graph datasets have no or unreliable ground-truth explanations for a given task. Here, we introduce a synthetic graph data generator, ShapeGGen, which can generate a variety of benchmark datasets (e.g., varying graph sizes, degree distributions, homophilic vs. heterophilic graphs) accompanied by ground-truth explanations. Further, the flexibility to generate diverse synthetic datasets and corresponding ground-truth explanations allows us to mimic the data generated by various real-world applications. We include ShapeGGen and several real-world graph datasets into an open-source graph explainability library, GraphXAI. In addition to synthetic and real-world graph datasets with ground-truth explanations, GraphXAI provides data loaders, data processing functions, visualizers, GNN model implementations, and evaluation metrics to benchmark the performance of GNN explainability methods.
Modern AI models contain much of human knowledge, yet understanding of their
internal representation of this knowledge remains elusive. Characterizing the
structure and properties of this representation will lead to improvements in
model capabilities and development of effective safeguards. Building on recent
advances in feature learning, we develop an effective, scalable approach for
extracting linear representations of general concepts in large-scale AI models
(language models, vision-language models, and reasoning models). We show how
these representations enable model steering, through which we expose
vulnerabilities, mitigate misaligned behaviors, and improve model capabilities.
Additionally, we demonstrate that concept representations are remarkably
transferable across human languages and combinable to enable multi-concept
steering. Through quantitative analysis across hundreds of concepts, we find
that newer, larger models are more steerable and steering can improve model
capabilities beyond standard prompting. We show how concept representations are
effective for monitoring misaligned content (hallucinations, toxic content). We
demonstrate that predictive models built using concept representations are more
accurate for monitoring misaligned content than using models that judge outputs
directly. Together, our results illustrate the power of using internal
representations to map the knowledge in AI models, advance AI safety, and
improve model capabilities.
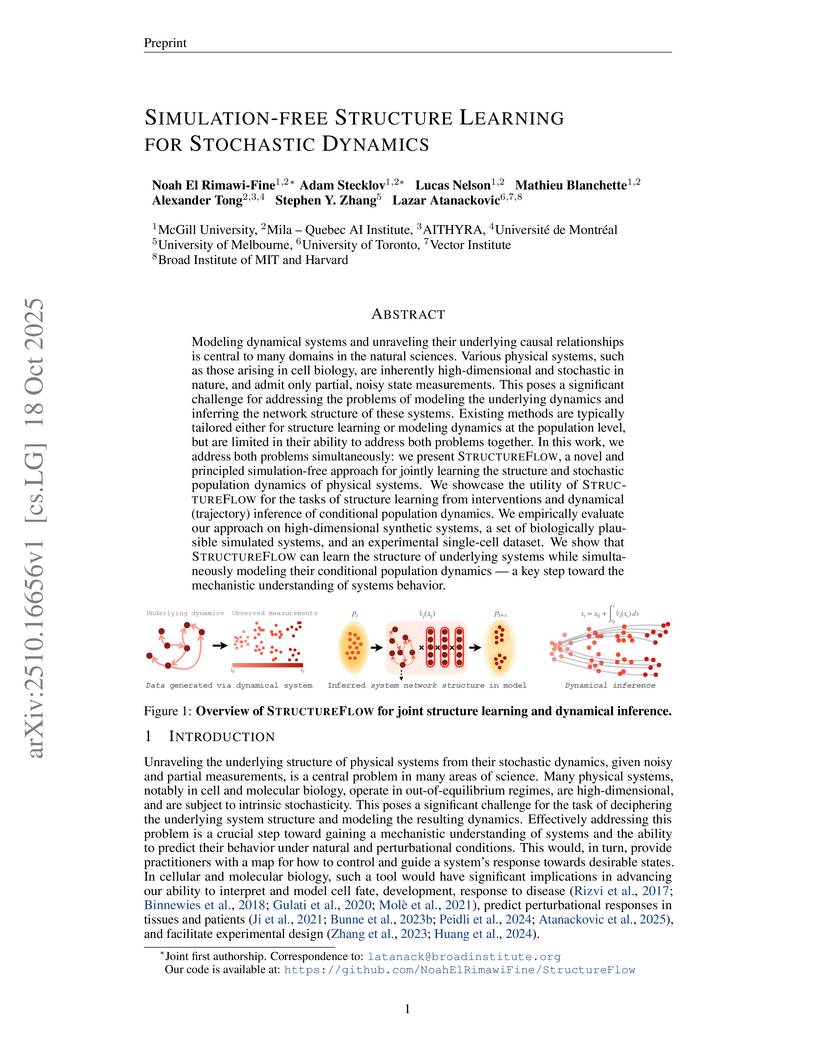
STRUCTUREFLOW presents a simulation-free framework for jointly inferring network structure and stochastic population dynamics from snapshot data, utilizing Schrödinger Bridges and score/flow matching. The method achieves superior structure learning and dynamical inference across synthetic and diverse biological systems, demonstrating strong generalization to unseen interventions in single-cell transcriptomics.
We study the problem active sequential hypothesis testing, also known as pure exploration: given a new task, the learner adaptively collects data from the environment to efficiently determine an underlying correct hypothesis. A classical instance of this problem is the task of identifying the best arm in a multi-armed bandit problem (a.k.a. BAI, Best-Arm Identification), where actions index hypotheses. Another important case is generalized search, a problem of determining the correct label through a sequence of strategically selected queries that indirectly reveal information about the label. In this work, we introduce In-Context Pure Exploration (ICPE), which meta-trains Transformers to map observation histories to query actions and a predicted hypothesis, yielding a model that transfers in-context. At inference time, ICPE actively gathers evidence on new tasks and infers the true hypothesis without parameter updates. Across deterministic, stochastic, and structured benchmarks, including BAI and generalized search, ICPE is competitive with adaptive baselines while requiring no explicit modeling of information structure. Our results support Transformers as practical architectures for general sequential testing.
Researchers from MBZUAI, Carnegie Mellon University, and Broad Institute developed a causal representation learning framework capable of achieving component-wise identifiability of latent variables from multimodal biomedical observations. The method recovers individual latent causal factors and their inter-modal relationships in a nonparametric setting, demonstrating consistency with established biomedical knowledge on a human phenotype dataset.
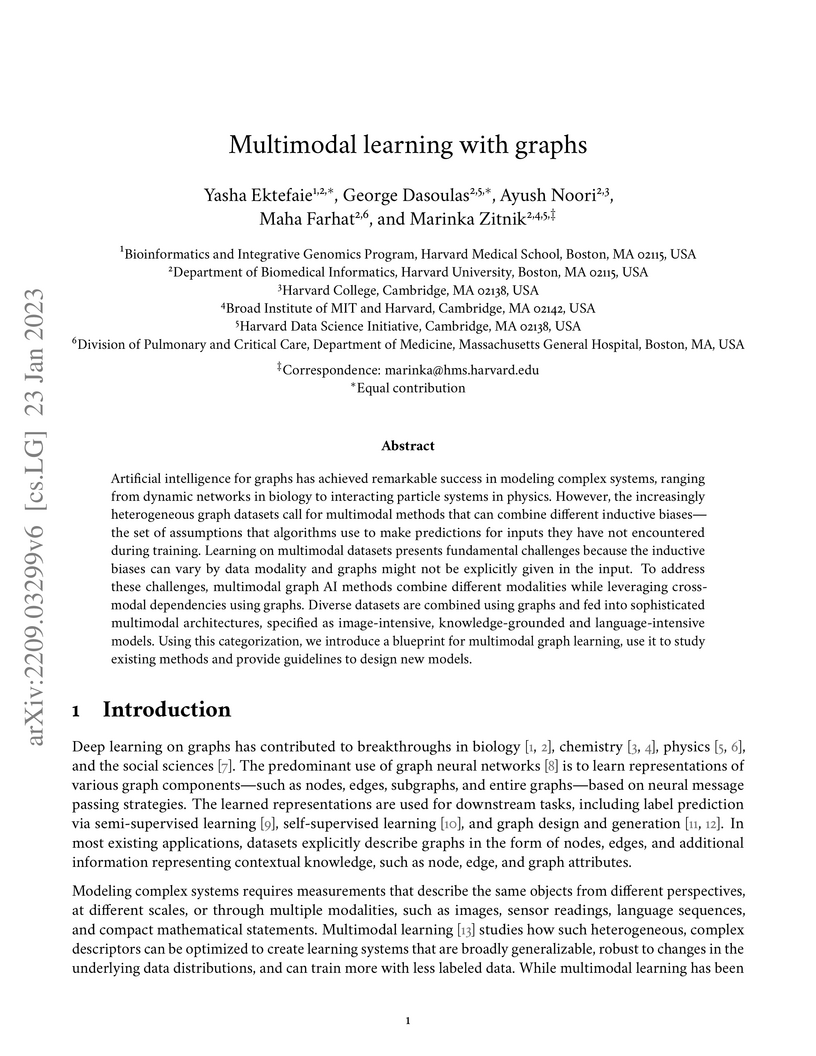
Artificial intelligence for graphs has achieved remarkable success in
modeling complex systems, ranging from dynamic networks in biology to
interacting particle systems in physics. However, the increasingly
heterogeneous graph datasets call for multimodal methods that can combine
different inductive biases: the set of assumptions that algorithms use to make
predictions for inputs they have not encountered during training. Learning on
multimodal datasets presents fundamental challenges because the inductive
biases can vary by data modality and graphs might not be explicitly given in
the input. To address these challenges, multimodal graph AI methods combine
different modalities while leveraging cross-modal dependencies using graphs.
Diverse datasets are combined using graphs and fed into sophisticated
multimodal architectures, specified as image-intensive, knowledge-grounded and
language-intensive models. Using this categorization, we introduce a blueprint
for multimodal graph learning, use it to study existing methods and provide
guidelines to design new models.
There are no more papers matching your filters at the moment.